| "Computer Science is no more about computers than astronomy is about telescopes." Edsger Dijkstra | "C'est aux algorithmes du monde vivant que s'intéresse aujourd'hui la biologie." François Jacob |
| "Le contraire de la recherche fondamentale ce n'est pas la recherche appliquée, c'est la recherche superficielle." Gilles Kahn |
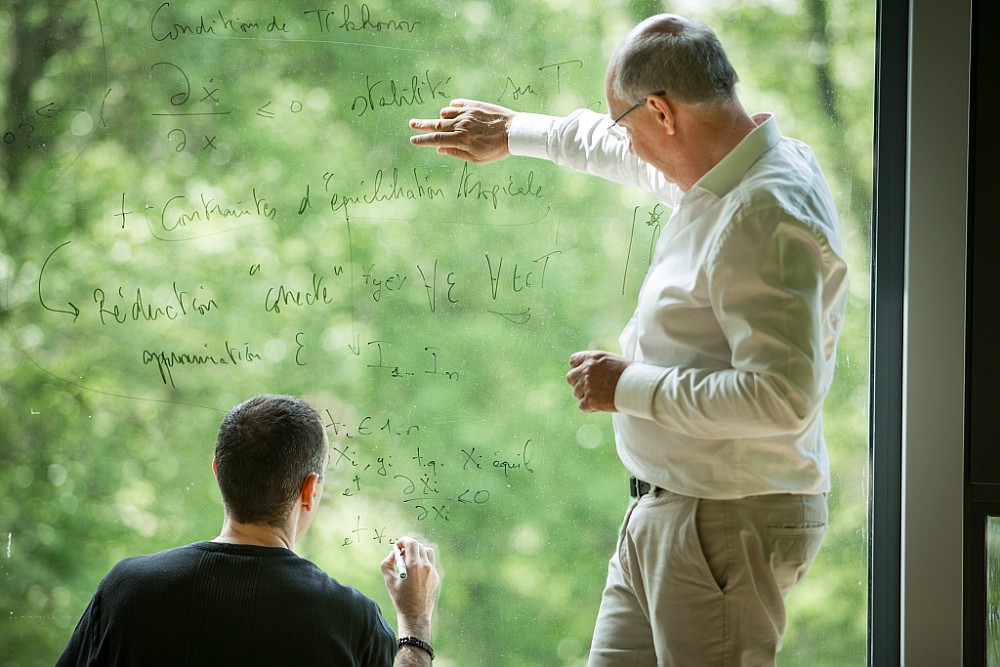
The Lifeware research team
Lifeware is a project-team of Inria, located in the Inria Saclay Center (formerly in Paris-Rocquencourt until January 2016) on the campus of Ecole Polytechnique in Palaiseau (and at Pasteur Institute until the creation of EPC Inbio at Inria Paris in January 2020).
Created in January 2014, as a follow-up of the Contraintes project-team, Lifeware aims at developing formal methods for understanding the cell machinery and establishing computational paradigms in cell biology. It is based on the vision of cells as machines, biochemical reaction networks as programs, and on the use of concepts and tools from computer science to master the complexity of cell processes.
Because of the importance of optimization problems in our research, we keep some research and teaching activity on Constraint Logic Programming for computing with partial information systems and practically solving NP-hard problems.
This project addresses fundamental research issues in computer science on the interplay between structure and dynamics in chemical reaction networks, and on their automated design. We contribute to a computational theory of chemical reaction networks (CRN), and develop since 2002 a CRN modelling, analysis and synthesis software, the Biochemical Abstract Machine, BIOCHAM (see also Inria Saclay showroom). The reaction rule-based language of this system allows us to reason about CRNs at different levels of abstraction, in their stochastic, differential, discrete, Boolean and graphical interpretations. We develop a variety of static analysis methods before going to simulations and dynamical analyses. We use quantitative temporal logics as a mean to formalise biological behaviours with imprecise data and to constrain CRN model building.
A tight integration between dry lab and wet lab efforts is also essential for the success of the project. This is achieved through long-term collaborations with biologists, pharmacologists and physicians who deal with advanced robotic, microfluidic and synthetic biology technologies to perform accurate observations at "single cell" level, and also to implement CRN programs in artificial devices for medical purposes.
Collaborations with top international leaders are established, and consolidated with student exchange programs, especially in the framework of the Institut Polytechnique de Paris.
Highlights
Paper published in Mathematical BIosciences
Ballif, Guillaume, Pfeiffer, Laurent, Ruess, Jakob. A partition method for bounding continuous-time Markov chain models of general reaction network. Mathematical Biosciences, 395, 2026. [ preprint ]
The place to be: our favourite conference CMSB we organised this year in Lyon CMSB 2025
next year CMSB 2026 will be associated to FLOC 2026, LIsbon, Portugal, July 2026.
François Fages recipient of the Skolem award of the 30th CADE conference 2025
Test of time award for his paperFrançois Fages. Associative-Commutative Unification. In Proc. of seventh Conf. on Automated Deduction CADE'84, volume 170 of LNCS. Springer-Verlag, 1984. [ preprint ]
La thèse d'Alexandre Tan -Lhernould en 180s
c'est sur https://www.youtube.com/watch?v=rotHmlgq9DU&t=91s à 38mn7s
Computational Systems Biology results based on Constraint Logic Programming
Fages, François, Hemery, Mathieu, Soliman, Sylvain. On BIOCHAM Symbolic Computation Pipeline for Compiling Mathematical Functions into Biochemistry. ACM Commun. Comput. Algebra, 58(2):15–22, 2025. [ preprint ]
Aghakhani, Sahar, Niarakis, Anna, Soliman, Sylvain. MetaLo: metabolic analysis of Logical models extracted from molecular interaction maps. Journal of Integrative Bioinformatics, 21(1):20230048, 2024. [ preprint ]
Crisci, Emma, Mahout, Maxime, Peres, Sabine. Computing Thermodynamically Consistent Elementary Flux Modes with Answer Set Programming. In Computational Methods in Systems Biology, 22nd Int. Conference, CMSB 2024, Pisa, Italy, September 16–18, 2024, Proceedings, pages 80–88, volume LNCS-14971 of Lecture Notes in Computer Science. Springer Nature Switzerland, 2024. [ preprint ]
Trinh, Giang, Benhamou, Belaid, Pastva, Samuel and Soliman, Sylvain. Scalable Enumeration of Trap Spaces in Boolean Networks via Answer Set Programming. In AAAI 2024 - The 38th Annual Conf. on Artificial Intelligence, pages 10714–10722, volume 38 of , 2024. [ preprint ]
Back to Logic Programming future
François Fages. A Constraint-based Mathematical Modeling Library in Prolog with Answer Constraint Semantics. In 17th Int. Symposium on Functional and Logic Programming, FLOPS 2024, pages 135–150, volume 14659 of LNCS. Springer-Verlag, 2024. [ preprint ]
Fages, François. On Teaching Constraint-based Modeling and Algorithms for Decision Support in Prolog. In Worskhop Proc. of the 40th Int. Conf. on Logic Programming, ICLP-WS 2024., 2024. [ preprint ]
Trinh, Van-Giang, Benhamou, Belaid, Soliman, Sylvain, Fages, François. Graphical conditions for the existence, unicity and number of regular models. In ICLP 2024 - 40th Int. Conf. on Logic Programming, 2024. [ preprint ]
Workshop for the HDR defense of Jakob Ruess on March 18-19th 2024
Three articles in TCS's special issue on Foundational Methods in Systems Biology:
Hemery, Mathieu, Fages, François. On a model of online analog computation in the cell with absolute functional robustness: algebraic characterization, function compiler and error control. Theoretical Computer Science, 991:114432, 2024. [ preprint ]
Trinh, Van-Giang, Benhamou, Belaid, Soliman, Sylvain. Trap spaces of Boolean networks are conflict-free siphons of their Petri net encoding. Theoretical Computer Science, 971:114073, 2023. [ preprint ]
Thibault Greugny, Eléa, Fages, François and Radulescu, Ovidiu, Szmolyan, Peter, Stamatas, Georgios. A Skin Microbiome Model with AMP interactions and Analysis of Quasi-Stability vs Stability in Population Dynamics. Theoretical Computer Science, 983:114294, 2023. [ preprint ]
Paper in Journal of Chemical Physics
Ruess, Jakob, Ballif, Guillaume, Aditya, Chetan. Stochastic chemical kinetics of cell fate decision systems: from single cells to populations and back. Journal of Chemical Physics, 159(18), 2023. [ preprint ]
Marine Collery PhD thesis public defence, October 12th 2023, 2pm, Amphi Sophie Germain, Build. Alan Turing, Inria Saclay
Expressive classification rule learning with an emphasis on learning from sequential data.
Sahar Aghakhani PhD thesis public defence, September 22nd 2023 2pm, G2 Building, CEA Évry
Computational modeling of metabolic reprogramming in rheumatoid arthritis synovial fibroblasts and cancer-associated fibroblasts
Back to Constraint Programming conference
Trinh, Van-Giang, Benhamou, Belaid, Soliman, Sylvain. Efficient Enumeration of Fixed Points in Complex Boolean Networks Using Answer Set Programming. In 29th Int. Conf. on Principles and Practice of Constraint Programming (CP 2023), pages 35:1–35:19, volume 280 of . Schloss Dagstuhl - Leibniz-Zentrum für Informatik, 2023. [ preprint ]
PhD graduation ceremony at Institut Polytechnique de Paris
to which we contributed for 3/251 PhD theses in 2022 with Dr Eléa Thibault-Greugny, Dr Jeremy Grignard and Dr Julien Martinelli
Three weeks meeting on Building Immune Digital Twin co-organized by Anna Niaraki
a fantastic exchange forum between life sciences and computational science at Institut Pascal, University Paris Saclay.
A paper presented at ICLR 2023
Collery, Marine, Bonnard, Philippe, Fages, François, Kusters, Remy. Neural-based classification rule learning for sequential data. In ICLR 2023 - The Eleventh Int. Conf. on Learning Representations, 2023. [ preprint ]
Mathieu Hemery at SIAM Computational Science in Engineering
presenting our approach to quadratization problems and their implementation in BIOCHAM
Jakob Ruess back to Lifeware !
where Jakob will develop his ERC starting grant BridgingScales project, welcome back !
Prolog 50th anniversary Symposium Paris Nov 10th 2022 small video
Document on the history of ISO Standard of Prolog to which our ancestor Inria project-teams Loco and Contraintes participated at Inria Rocquencourt.
As an example of current use of Prolog, it is worth noting that our cutting-edge CRN modelling, analysis and synthesis software BIOCHAM is implemented in Prolog, for several good reasons (rapid prototyping, constraint solving libraries, symbolic computation, easy parsing and syntax tree manipulation).
Paper published in PLOS computational biology
Aghakhani, Sahar, Soliman, Sylvain, Niarakis, Anna. Metabolic Reprogramming in Rheumatoid Arthritis Synovial Fibroblasts: a Hybrid Modeling Approach. PLoS Computational Biology, 18(12):e1010408, 2022. [ preprint ]
Elea Thibault-Greugny PhD thesis public defence, October 4th 2022 2pm,
Elea Thibault-Greugny. Computational modeling approaches to multifactorial aspects of atopic dermatitis. PhD Thesis, Institut Polytechnique de Paris, 2022. [ preprint ]
3 papers accepted at our favourite conference CMSB
Best Paper Award given to the first:Mathieu Hemery, François Fages. Algebraic Biochemistry: a Framework for Analog Online Computation in Cells. In CMSB'22: Proc. of the twentieth international Conf. on Computational Methods in Systems Biology, volume 13447 of LNCS. Springer-Verlag, 2022. [ preprint ] [ slides ] [ video ]
Eléa Thibault Greugny, Georgios N Stamatas and François Fages. Stability versus Meta-stability in a Skin Microbiome Model. In CMSB'22: Proc. of the twentieth international Conf. on Computational Methods in Systems Biology, volume 13447 of LNCS. Springer-Verlag, 2022. [ preprint ] [ slides ]
Van-Giang Trinh, Belaid Benhamou, Kunihiko Hiraishi, Sylvain Soliman. Minimal trap spaces of Logical models are maximal siphons of their Petri net encoding. In CMSB'22: Proc. of the twentieth international Conf. on Computational Methods in Systems Biology, volume 13447 of LNCS. Springer-Verlag, 2022. [ preprint ]
Jeremy Grignard PhD thesis public defence, June 17th 2022 2pm
Jeremy Grignard. Méthodes computationnelles pour améliorer les phases primaires de recherche de nouveaux médicaments. PhD Thesis, Institut Polytechnique de Paris, 2022. [ preprint ]
A paper with Servier accepted at PLOS Computational Biology
Jeremy Grignard, Véronique Lamamy, Eva Vermersch, Philippe Delagrange, Jean-Philippe Stephan, Thierry Dorval, François Fages. Mathematical modeling of the microtubule detyrosination/tyrosination cycle for cell-based drug screening design. PLOS Computational Biology, 18(6), 2022.
Julien Martinelli PhD thesis defense, Feb 18th 2022
Julien Martinelli. On learning mechanistic models from time series data with applications to personalised chronotherapies. PhD Thesis, Institut Polytechnique de Paris, 2022. [ preprint ]
Jeremy Grignard on France Culture, Jan12th 2021.
Genomique et IA (from 40mn48s)
The COVID19 disease map: a Bioinformatics feat ! published in Mol. Sys. Biol.
Ostaszewski, Marek, Niarakis, Anna, Mazein, Alexander, Kuperstein, Inna, Phair, Robert and Orta-Resendiz, Aurelio, Singh, Vidisha, Aghamiri, Sara Sadat, Acencio, Marcio Luis et al. COVID19 Disease Map, a computational knowledge repository of virus-host interaction mechanisms. Molecular Systems Biology, 17(10):e10387, 2021. [ preprint ]
An article presented at ECCB 2021, published in Bioinformatics
Julien Martinelli, Sandrine Dulong, Xiao-Mei Li , Michèle Teboul, Sylvain Soliman, Francis Lévi, François Fages, Annabelle Ballesta. Model learning to identify systemic regulators of the peripheral circadian clock. Bioinformatics, 37:i401-i409, 2021. [ preprint ] [ slides ] [ video ]
A paper accepted at CMSB 2021 and CASC 2021
Mathieu Hemery, François Fages, Sylvain Soliman. Compiling Elementary Mathematical Functions into Finite Chemical Reaction Networks via a Polynomialization Algorithm for ODEs. In CMSB'21: Proc. of the nineteenth international Conf. on Computational Methods in Systems Biology, volume 12881 of LNCS. Springer-Verlag, 2021. [ preprint ]
A paper published in Cancers
Aghakhani, Sahar, Zerrouk, Naouel, Niarakis, Anna. Metabolic Reprogramming of Fibroblasts as Therapeutic Target in Rheumatoid Arthritis and Cancer: Deciphering Key Mechanisms Using Computational Systems Biology Approaches. Cancers, 2021. [ preprint ]
An article on SBML 3, the latest version of the Systems Biology Markup Language standard
Sarah M. Keating, Dagmar Waltemath, Matthias König, Fengkai Zhang, Andreas Dräger and Claudine Chaouiya, Frank T. Bergmann, Andrew Finney , Colin S. Gillespie et al. SBML Level 3: an extensible format for the exchange and reuse of biological models. Molecular Systems Biology, 16(8):1–21, 2020. [ preprint ]
A paper published in Bioinformatics
Aghamiri, Sara Sadat, Singh, Vidisha, Naldi, Aurélien, Helikar, Tomas, Soliman, Sylvain and Niarakis, Anna. Automated inference of Boolean models from molecular interaction maps using CaSQ. Bioinformatics, 2020. [ preprint ]
Two papers accepted at CMSB 2020
Mathieu Hemery, François Fages, Sylvain Soliman. On the Complexity of Quadratization for Polynomial Differential Equations. In CMSB'20: Proc. of the eighteenth international Conf. on Computational Methods in Systems Biology, LNCS. Springer-Verlag, 2020. [ preprint ]
Elisabeth Degrand, François Fages, Sylvain Soliman. Graphical Conditions for Rate Independence in Chemical Reaction Networks. In CMSB'20: Proc. of the eighteenth international Conf. on Computational Methods in Systems Biology, volume 12314 of LNCS. Springer-Verlag, 2020. [ preprint ] [ slides ] [ video ]
Biomachines for real
Francisco Santos Schneider, Patrick Amar, Asma Bahri, Julien Espeut, Julie Baptiste, Mellis Alali, François Fages, Franck Molina. Biomachines for Medical Diagnosis. Advanced Materials Letters, 11(4):1535–1558, 2020.
Book chapter on Biomachines
Fages, François, Molina, Franck. The Cell, a Chemical Analog Calculator. In Symbolic Approaches to Modeling and Analysis of Biological Systems, pages 235–254. John Wiley Sons, Ltd, 2023. [ preprint ]
François Fages, Franck Molina. La cellule, un calculateur analogique chimique. In Approches symboliques de la modélisation et de l'analyse des systèmes biologiques, pages 255–273. ISTE group, Ed Ltd, London, 2022. [ preprint ] [ video ]
Book chapter in an encyclopedia of Artificial Intelligence
François Fages. Artificial Intelligence in Biological Modeling. In A Guided Tour of Artificial Intelligence Research, Volume III: Interfaces and Applications of AI. Springer-Verlag, 2020. [ preprint ]
Spin-off Inbio
Inbio has now officially left our group to create a new research group of the Inria Paris research centre at Pasteur Institute
Two papers in PLOS Computational Biology from InBIo group
Cepeda-Humerez, Sarah Anhala, Ruess, Jakob and Tkavcik, Gavsper. Estimating information in time-varying signals. PLoS Computational Biology, 15(9):e1007290, 2019. [ preprint ]
Ruess, Jakob, Plevska, Marovs, Guet, Cvalin C, Tkavcik, Gavsper. Molecular noise of innate immunity shapes bacteria-phage ecologies. PLoS Computational Biology, 2019. [ preprint ]
Prix La Recherche 2019 
 F. Fages with O. Bournez LIX next floor
F. Fages with O. Bournez LIX next floor
We are very honoured to receive the 2019 Prix La Recherche - science de l'information for our article (best paper award CMSB 2017) [slides]:
Fages, François, Le Guludec, Guillaume and Bournez, Olivier, Pouly, Amaury. Strong Turing Completeness of Continuous Chemical Reaction Networks and Compilation of Mixed Analog-Digital Programs. In CMSB'17: Proc. of the fiveteen international Conf. on Computational Methods in Systems Biology, pages 108–127, volume 10545 of LNCS. Springer-Verlag, 2017. [ preprint ]
A paper accepted in Science Advances
Meredith, Hannah, Andreani, Virgile, Ma, Helena , Lopatkin, Allison, Lee, Anna, Anderson, Deverick, Batt, Gregory, You, Lingchong. Applying ecological resistance and resilience to dissect bacterial antibiotic responses. Science Advances, 4(12):eaau1873, 2018. [ preprint ]
A paper published in the Journal of Theoretical Biology
Adrien Baudier, François Fages, Sylvain Soliman. Graphical Requirements for Multistationarity in Reaction Networks and their Verification in BioModels. Journal of Theoretical Biology, 459:79–89, 2018. [ preprint ]
A paper in cover page of Molecular Systems Biology
Alexis Courbet, Patrick Amar, François Fages, Eric Renard, Franck Molina. Computer-aided biochemical programming of synthetic microreactors as diagnostic devices. Molecular Systems Biology, 14(4), 2018. [ preprint ]
A paper in Scientific Reports
Lugagne, Jean-Baptiste, Jain, Srajan, Ivanovitch, Pierre, Ben Meriem, Zacchary, Vulin, Clément and Fracassi, Chiara, Batt, Grégory, Hersen, Pascal. Identification of individual cells from z-stacks of bright-field microscopy images. Scientific Reports, 8(1):11455, 2018.
A paper accepted in IEEE/ACM Transactions on Computational Biology and Bioinformatics
François Fages, Thierry Martinez, David Rosenblueth, Sylvain Soliman. Influence Networks compared with Reaction Networks: Semantics, Expressivity and Attractors. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 2018. [ preprint ]
Two papers published in the same issue of Nature Communications !
Remy Chait, Jakob Ruess, Tobias Bergmiller and Gavsper Tkavcik, Cvalin Guet. Shaping bacterial population behavior through computer-interfaced control of individual cells. Nature Communications, 8(1):1535, 2017.
Jean-Baptiste Lugagne, Sebastian Sosa Carrillo and Melanie Kirch, Agnes Köhler, Gregory Batt and Pascal Hersen. Balancing a genetic toggle switch by real-time feedback control and periodic forcing. Nature Communications, 8(1):1671, 2017.
BIOCHAM v4.0 released
complete rewriting of BIOCHAM with online notebooks, short tutorial and historical tutorial
A paper accepted in Bioinformatics
Palaniappan, Sucheendra K., Bertaux, François , Pichené, Matthieu, Fabre, Eric, Batt, Gregory, Genest, Blaise. Abstracting the dynamics of biological pathways using information theory: a case study of apoptosis pathway. Bioinformatics, 33(13):1980–1986, 2017. [ preprint ]
A paper accepted in J. Chemical Physics
Jakob Ruess, Heinz Koeppl, Christoph Zechner. Sensitivity estimation for stochastic models of biochemical reaction networks in the presence of extrinsic variability. The Journal of Chemical Physics, 146(12):124122, 2017.
Best Paper Award at CMSB 2017 and second paper
The first one solving a long standing open problem in chemical reaction network theory
Fages, François, Le Guludec, Guillaume and Bournez, Olivier, Pouly, Amaury. Strong Turing Completeness of Continuous Chemical Reaction Networks and Compilation of Mixed Analog-Digital Programs. In CMSB'17: Proc. of the fiveteen international Conf. on Computational Methods in Systems Biology, pages 108–127, volume 10545 of LNCS. Springer-Verlag, 2017. [ preprint ]
Carcano, Arthur, Fages, François, Soliman, Sylvain. Probably Approximately Correct Learning of Regulatory Networks from Time-Series Data. In CMSB'17: Proc. of the fifteenth international Conf. on Computational Methods in Systems Biology, pages 74–90, volume 10545 of , 2017. [ preprint ]
InBio starts!
InBio is an Inria/Pasteur research group created in Feb 2017 and headed by Grégory Batt.
CellStar paper published in Royal Society Interface
Cristian Versari, Szymon Stoma, Kirill Batmatov , Artémis Llamosi, Filip Moroz, Adam Kaczmarek , Matt Deyell, Cédric Lhoussaine, P. Hersen and G. Batt. Long-term tracking of budding yeast cells in brightfield microscopy: CellStar and the Evaluation Platform. Royal Society Interface, 14, 2017.
PhD Thesis of Jean-Baptiste Lugagne
13 Dec 2016 in Paris
HDR of Sylvain Soliman
7 Dec 2016 in Palaiseau
A paper accepted in Biosystems
Pauline Traynard, Céline Feillet, Sylvain Soliman, Franck Delaunay, François Fages. Model-based Investigation of the Circadian Clock and Cell Cycle Coupling in Mouse Embryonic Fibroblasts: Prediction of RevErb-alpha Up-Regulation during Mitosis. Biosystems, 149:59–69, 2016. [ preprint ]
A paper accepted at ECCB'16
Pauline Traynard, Adrien Fauré, François Fages, Denis Thieffry. Logical model specification aided by model- checking techniques: application to the mammalian cell cycle regulation. Bioinformatics, 32(17):i772-i780, 2016. [ preprint ]
A paper accepted at CMSB'16
François Fages, Thierry Martinez, David Rosenblueth, Sylvain Soliman. Influence Systems vs Reaction Systems. In CMSB'16: Proc. of the fourteenth international Conf. on Computational Methods in Systems Biology, pages 98–115, volume 9859 of LNCS. Springer-Verlag, 2016. [ preprint ]
PhD Thesis defense of François Bertaux
Wednesday June 15th at Inria Paris.
A paper accepted in PLoS Computational Biology
A. Llamosi, A.M. Gonzales-Vargas, E. Cinquemani , G. Ferrari-Trecate, P. Hersen, G. Batt. What Population Reveals about Individual Cell Identity: Single-Cell Parameter Estimation of Models of Gene Expression in Yeast. PLOS Computational Biology, 9(12), 2016.
PhD Thesis defense of Pauline Traynard
Tuesday May 10th at ENS.
PhD Theses season
Artemis Llamosi on Tuesday 15 December and Thierry Martinez on Thursday 17 December will defend their PhD thesis !
A paper accepted in PLoS ONE
Tai-Yin Chiu, Hui-Ju K. Chiang, Ruei-Yang Huang , Jie-Hong R. Jiang, François Fages. Synthesizing Configurable Biochemical Implementation of Linear Systems from Their Transfer Function Specifications. PLoS ONE, 10(9):e0137442, 2015.
PhD Defense of Steven Gay
May 26, 2015, University Paris-DIderot.
A paper accepted in ACM TOMACS
Hui-Ju Chiang, François Fages, Jie-Hong Jiang, Sylvain Soliman. Hybrid Simulations of Heterogeneous Biochemical Models in SBML. ACM Transactions on Modeling and Computer Simulation (TOMACS), 25(2):14:1–14:22, 2015. [ preprint ]
A paper accepted in Constraints
Faten Nabli, Thierry Martinez, François Fages, Sylvain Soliman. On Enumerating Minimal Siphons in Petri nets using CLP and SAT solvers: Theoretical and Practical Complexity. Constraints, 21(2):251–276, 2016. [ preprint ]
Xavier Duportet laureate of Doctors-Entrepreneurs awards
for his PhageX project, the antibiotics of the future, congrats !!!
A paper accepted in PLoS Computational Biology
F. Bertaux, S. Stoma, D. Drasdo, G. Batt. Modeling dynamics of cell-to-cell variability in TRAIL-induced apoptosis explains fractional killing and predicts reversible resistance. PLoS Computational Biology, 10(10):e1003893, 2014.
A paper accepted in Algorihms for Molecular Biology
Sylvain Soliman, François Fages, Ovidiu Radulescu. A constraint solving approach to model reduction by tropical equilibration. Algorithms for Molecular Biology, 9(24), 2014.
PhD Defense of David Fournier
November 27th 2014, University Paris-Diderot
Best student paper prize at CMSB'14, congratulations Pauline !
Pauline Traynard, François Fages, Sylvain Soliman. Trace Simplifications preserving Temporal Logic Formulae with Case Study in a Coupled Model of the Cell Cycle and the Circadian Clock. In CMSB'14: Proc. of the twelth international Conf. on Computational Methods in Systems Biology, pages 114–128, LNCS. Springer-Verlag, 2014. [ preprint ] [ slides ]
Exceptional PhD Defense of Xavier Duportet
November 14th 2014, Pasteur Institute
Release of Biocham 3.6
New release of Biocham with new features for hybrid simulation, improved SBML-3 interface and new version of biocham-web
Keynote talk at ECAI'14
Slides (pdf, pptx), see also invited talk paper at ICDCIT'14:
François Fages. Cells as Machines: towards Deciphering Biochemical Programs in the Cell (Invited Talk). In Proc. 10th Int. Conf. on Distributed Computing and Internet Technology ICDCIT'14, pages 50–67, volume 8337 of Lecture Notes in Computer Science. Springer-Verlag, 2014. [ preprint ] [ video ]
A paper accepted in TCS
François Fages, Steven Gay, Sylvain Soliman. Inferring Reaction Systems from Ordinary Differential Equations. Theoretical Computer Science, 599:64–78, 2015.
Laureate of the French Academy of Sciences
François Fages is very honoured to receive the Michel Monpetit prize 2014 of the French Academy of Sciences for his contributions to fundamental computer science (unification theory and constraint logic programming) and computational systems biology (modeling of biochemical networks and design and supervision of the implementation of the BIOCHAM software).
Slides (pdf, pptx) and talk in French at College de France.
A paper accepted in Discrete Applied Mathematics
Steven Gay, François Fages, Thierry Martinez, Sylvain Soliman, Christine Solnon. On the subgraph Epimorphism Problem. Discrete Applied Mathematics, 162:214–228, 2014. [ preprint ]
HDR defense of Grégory Batt
Grégory Batt brightly defended his ''Habilitation à Diriger des Recherches" (manuscript, slides,)
An article in the Bulletin of Mathematical Biology
refining the 10 years old result of C. Soulé on feedback circuits and multistationarity
Sylvain Soliman. A stronger necessary condition for the multistationarity of chemical reaction networks. Bulletin of Mathematical Biology, 75(11):2289–2303, 2013. [ preprint ]
A paper accepted in PLoS Computational Biology
Szymon Stoma, Alexandre Donzé, François Bertaux, Oded Maler, Grégory Batt. STL-based analysis of TRAIL-induced apoptosis challenges the notion of type I/type II cell line classification. PLoS Computational Biology, 9(5):e1003056, 2013.
PhD Defense of Faten Nabli
July 10th 2013, Univ. Paris-Diderot
Release of Biocham-web in May 2013
Biocham-Web provides you access to the latest version of Biocham as a web service.
A paper accepted at CP 2012
Faten Nabli, François Fages, Thierry Martinez, Sylvain Soliman. A Boolean Model for Enumerating Minimal Siphons and Traps in Petri-nets. In Proc. of CP'2012, 18th Int. Conf. on Principles and Practice of Constraint Programming, pages 798–814, volume 7514 of LNCS. Springer-Verlag, 2012. [ preprint ] [ slides ] [ poster ]
A paper accepted in PNAS
Jannis Uhlendorf, Agnés Miermont, Thierry Delaveau, Gilles Charvin, François Fages and Samuel Bottani, Gregory Batt, Pascal Hersen. Long-term model predictive control of gene expression at the population and single-cell levels. Proc. of the National Academy of Sciences USA, 109(35):14271–14276, 2012.
A paper accepted in MolSysBiol, with Robert Lefkowitz, Nobel Prize in Chemistry 2012
Domitille Heitzler, Guillaume Durand, Nathalie Gallay, Aurélien Rizk, Seungkirl Ahn, Jihee Kim, Jonathan D. Violin, Laurence Dupuy and Christophe Gauthier, Vincent Piketty, Pascale Crépieux, Anne Poupon, Frédérique Clément, François Fages, Robert J. Lefkowitz, Eric Reiter. Competing G protein-coupled receptor kinases balance G protein and β-arrestin signaling. Molecular Systems Biology, 8(590), 2012.
A paper accepted in Algorithms for Molecular Biology
Sylvain Soliman. Invariants and Other Structural Properties of Biochemical Models as a Constraint Satisfaction Problem. Algorithms for Molecular Biology, 7(15), 2012.
Release of FO-CTL(ℝlin) in January 2012
FO-CTL(ℝlin) is a constraint solver for full First-Order Computation Tree Logic formulae with linear arithmetic over the reals in constrained transition systems (CTS). CTS are transition systems where both states and transitions are described with constraints.
Two papers accepted in TCS
Aurélien Rizk, Grégory Batt, François Fages, Sylvain Soliman. Continuous Valuations of Temporal Logic Specifications with applications to Parameter Optimization and Robustness Measures. Theoretical Computer Science, 412(26):2827–2839, 2011. [ preprint ]
Elisabetta De Maria, François Fages, Aurélien Rizk, Sylvain Soliman. Design, Optimization, and Predictions of a Coupled Model of the Cell Cycle, Circadian Clock, DNA Repair System, Irinotecan Metabolism and Exposure Control under Temporal Logic Constraints. Theoretical Computer Science, 412(21):2108–2127, 2011. [ preprint ]
A paper accepted at PPDP 2010
Thierry Martinez. Semantics-preserving translations between Linear Concurrent Constraint Programming and Constraint Handling Rules. In Proc. of PPDP'10, Int. Conf. on Principles and Practice of Declarative Programming, Edinburgh, UK, pages 57–66. ACM, 2010. [ preprint ]
Release of Biocham 3 in June 2010
Biocham 3 is now available including a new SBGN graphical editor, new methods for model reduction, parameter optimization and robustness analyses w.r.t. temporal logic specifications.
A paper accepted at ECCB 2010 and in Bioinformatics
Steven Gay, Sylvain Soliman, François Fages. A Graphical Method for Reducing and Relating Models in Systems Biology. Bioinformatics, 26(18):i575–i581, 2010.
A paper accepted at CP 2009
François Fages, Aurélien Rizk. From Model-Checking to Temporal Logic Constraint Solving. In Proc. of CP'2009, 15th Int. Conf. on Principles and Practice of Constraint Programming, pages 319–334, LNCS. Springer-Verlag, 2009. [ preprint ]
A paper accepted in JTB
Sriram Krishnamachari, Sylvain Soliman, François Fages. Dynamics of the interlocked positive feedback loops explaining the robust epigenetic switching in Candida albicans. Journal of Theoretical Biology, 258(1):71–88, 2009. [ preprint ]
A paper accepted at ISMB 2009 and in Bioinformatics
Aurélien Rizk, Grégory Batt, François Fages, Sylvain Soliman. A general computational method for robustness analysis with applications to synthetic gene networks. Bioinformatics, 12(25):il69–il78, 2009.
Release of Rules2CP and PKML in April 2009
Rules2CP is a general purpose rule-based modeling language for constraint programming. It aims at making constraint programming technology easier to use by non-programmers, by modeling combinatorial optimization problems with logical rules and elementary data structures. The Packing Knowledge Modeling Language (PKML) is a Rules2CP library dedicated to industrial bin packing problems.
A paper accepted in TCS
François Fages, Aurélien Rizk. On Temporal Logic Constraint Solving for the Analysis of Numerical Data Time series. Theoretical Computer Science, 408(1):55–65, 2008. [ preprint ]
A paper in a book chapter
François Fages, Sylvain Soliman. Model Revision from Temporal Logic Properties in Systems Biology. In Probabilistic Inductive Logic Programming, pages 287–304, volume 4911 of LNCS. Springer-Verlag, 2008. [ preprint ]
A paper accepted in TCS
François Fages, Sylvain Soliman. Abstract Interpretation and Types for Systems Biology. Theoretical Computer Science, 403(1):52–70, 2008. [ preprint ]
Release of CHRat in July 2008
The CHRat language is a modular version of the Constraint Handling Rules language CHR, called CHRat for modular CHR with ask and tell. Any constraint defined in a CHRat component can be reused both in rules and guards in another CHRat component to define new constraint solvers.
Release of Biocham 2.7 in April 2008
Biocham 2.7 releases a powerful parameter search procedure with respect to temporal logic quantitative properties, a stochastic simulator and an inference system for conservation laws.
A tutorial paper at SFM 2008
François Fages, Sylvain Soliman. Formal Cell Biology in BIOCHAM. In 8th Int. School on Formal Methods for the Design of Computer, Communication and Software Systems: Computational Systems Biology SFM'08, pages 54–80, volume 5016 of LNCS. Springer-Verlag, 2008. [ preprint ]
A paper accepted at FMSB 2008
François Fages, Sylvain Soliman. From reaction models to influence graphs and back: a theorem. In Proc. of Formal Methods in Systems Biology FMSB'08, LNCS. Springer-Verlag, 2008. [ preprint ]
Grand Prize Finalist + First Place Foundational Research at iGEM 2007
For the first participation of a French team to the iGEM competition organised by the MIT on synthetic biology, the Paris team, with Aurélien Rizk using BIOCHAM, was Grand Prize Finalist plus First Place - Foundational Research (slides).