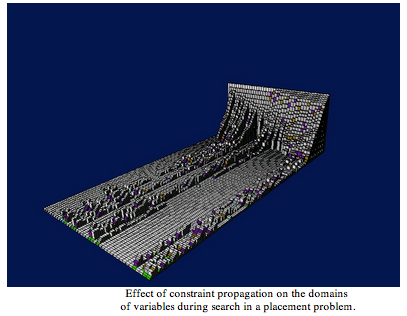
Software development
- "La simplicité est la sophistication suprême." Léonard de Vinci
 Biocham the Biochemical Abstract Machine for modeling, analyzing and synthesizing chemical reaction networks (CRN) in both perspectives of systems biology and synthetic biology. Biocham 4 is a complete rewriting of Biocham 3 for easier maintenance.
Biocham the Biochemical Abstract Machine for modeling, analyzing and synthesizing chemical reaction networks (CRN) in both perspectives of systems biology and synthetic biology. Biocham 4 is a complete rewriting of Biocham 3 for easier maintenance.
Biocham-4 includes
- a notebook on-line
- a short tutorial ( video with comments in French)
- an historical tutorial
- new features for
- compiling mathematical functions or programs in chemical reaction networks (CRN)
- influence systems with forces
- in addition to Biocham features for
- differential simulation, stochastic simulation, Boolean model-checking and Boolean attractors
- specifying semi-qualitative semi-quantitative behaviors in quantitative temporal logic
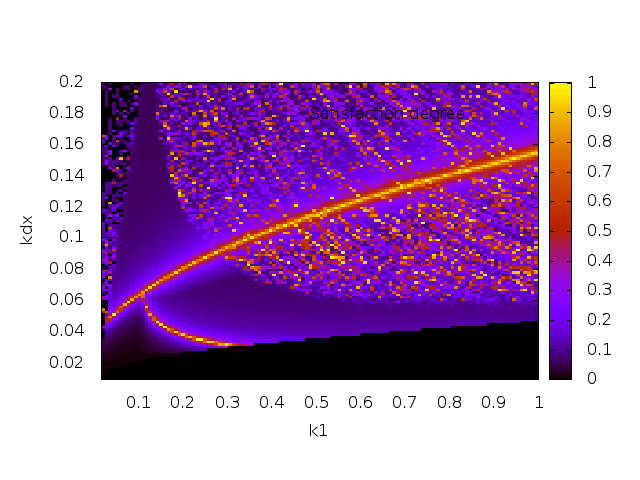
- verifying such behaviors, estimating parameter sensitivity and robustness
- searching and optimizing parameter values in high dimension
- detecting model reductions by subgraph epimorphisms
- various static analyses: dimension analysis, multi-stability, tropicalisation ...
-
Laurence Calzone, François Fages, Sylvain Soliman. BIOCHAM: An Environment for Modeling Biological Systems and Formalizing Experimental Knowledge. Bioinformatics, 22(14):1805–1807, 2006. [ preprint ]
-
François Fages, Sylvain Soliman. Formal Cell Biology in BIOCHAM. In 8th Int. School on Formal Methods for the Design of Computer, Communication and Software Systems: Computational Systems Biology SFM'08, pages 54–80, volume 5016 of LNCS. Springer-Verlag, 2008. [ preprint ]
 CaSQ: Celldesigner as Sbml-Qual CellDesigner model to SBML-Qual encoded logical model converter, co-developed by Anna Niarakis and Sylvain Soliman.
CaSQ: Celldesigner as Sbml-Qual CellDesigner model to SBML-Qual encoded logical model converter, co-developed by Anna Niarakis and Sylvain Soliman. Aghamiri, Sara Sadat, Singh, Vidisha, Naldi, Aurélien, Helikar, Tomas, Soliman, Sylvain and Niarakis, Anna. Automated inference of Boolean models from molecular interaction maps using CaSQ. Bioinformatics, 2020. [ preprint ]
Trappist a tool for computing minimal trap spaces of a Boolean model, developed by Sylvain Soliman.
Trinh, Van-Giang, Benhamou, Belaid, Soliman, Sylvain. Trap spaces of Boolean networks are conflict-free siphons of their Petri net encoding. Theoretical Computer Science, 971:114073, 2023. [ preprint ]
- MetaLo an opensource Python package that enables the coupling of Boolean models inferred from process description molecular interaction maps with generic core metabolic networks. Developed by Sylvain Soliman, Sahar Aghakhani and Anna Niarakis.
Aghakhani, Sahar, Niarakis, Anna, Soliman, Sylvain. MetaLo: metabolic analysis of Logical models extracted from molecular interaction maps. Journal of Integrative Bioinformatics, 21(1):20230048, 2024. [ preprint ]
- Pack modeling a SWI-Prolog pack for mathematical modellng with constraints on subscripted variables defined by set comprehension in Prolog, developed by François Fages.
François Fages. A Constraint-based Mathematical Modeling Library in Prolog with Answer Constraint Semantics. In 17th Int. Symposium on Functional and Logic Programming, FLOPS 2024, pages 135–150, volume 14659 of LNCS. Springer-Verlag, 2024. [ preprint ]
No longer maintained
- BIOCHAM 3.7 is a previous version of the Biochemical Abstract Machine software for modelilng and analysing biochemical reaction networks. BIOCHAM-web allows you to try it on-line through a web service. Beyond making simulations of different kinds, BIOCHAM provides unique features for
- importing and exporting biochemical reaction networks (CRNs) s in different formalisms and formats,
- performing various static analyses of CRNs,
- specifying behaviors in (quantitative) temporal logic,
- verifying tempoal properties by model-checking, and estimating parameter sensitivity and robustness,
- searching parameter values under temporal logic constraints.
- FO-CTL is a constraint solver for full First-Order Computation Tree Logic with linear arithmetic over the reals, FO-CTL(ℝlin), in Constrained Transition Systems.
- ClpZinc a Horn clause front-end for the MiniZInc modelling language for expressing search strategies by constraints
- MiniZinc-CMAES a stochastic optimization backend for the MiniZinc modelling language
- Rules2CP a general purpose rule-based modelling language for constraint solving.
- CLPGUI a generic graphical user interface for constraint logic programming.
Public data, models, notebooks and benchmarks related to publications.
- TCS 23
- Companion notebook to upload on BIOCHAM Jupyter notebook (alternatively see result in pdf,html or upload Biocham bc files)
-
Thibault Greugny, Eléa, Fages, François and Radulescu, Ovidiu, Szmolyan, Peter, Stamatas, Georgios. A Skin Microbiome Model with AMP interactions and Analysis of Quasi-Stability vs Stability in Population Dynamics. Theoretical Computer Science, 983:114294, 2023. [ preprint ]
- CMSB 2022
- Companion notebook to upload on BIOCHAM Jupyter notebook (alternatively see result in html or upload biocham bc files)
-
Mathieu Hemery, François Fages. Algebraic Biochemistry: a Framework for Analog Online Computation in Cells. In CMSB'22: Proc. of the twentieth international Conf. on Computational Methods in Systems Biology, volume 13447 of LNCS. Springer-Verlag, 2022. [ preprint ] [ slides ] [ video ]
- CMSB 2022b
- Companion notebook to upload on BIOCHAM Jupyter notebook (alternatively see result in pdf,html or upload Biocham bc files)
-
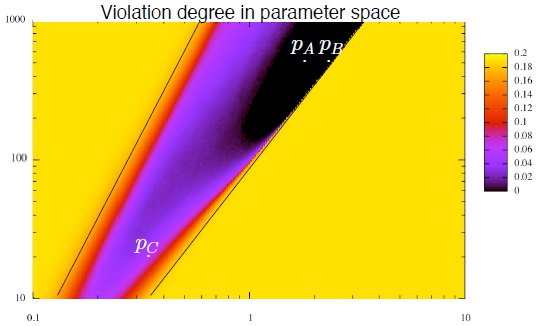
Eléa Thibault Greugny, Georgios N Stamatas and François Fages. Stability versus Meta-stability in a Skin Microbiome Model. In CMSB'22: Proc. of the twentieth international Conf. on Computational Methods in Systems Biology, volume 13447 of LNCS. Springer-Verlag, 2022. [ preprint ] [ slides ]
- CMSB 2021
- Companion notebook to save as local file and upload on BIOCHAM Jupyter notebook with v4.5.14 kernel
-
Mathieu Hemery, François Fages, Sylvain Soliman. Compiling Elementary Mathematical Functions into Finite Chemical Reaction Networks via a Polynomialization Algorithm for ODEs. In CMSB'21: Proc. of the nineteenth international Conf. on Computational Methods in Systems Biology, volume 12881 of LNCS. Springer-Verlag, 2021. [ preprint ]
- CMSB 2020a
- Benchmark (zip file) and BIOCHAM notebooks for minimizing either number of species or the number of reactions
-
Mathieu Hemery, François Fages, Sylvain Soliman. On the Complexity of Quadratization for Polynomial Differential Equations. In CMSB'20: Proc. of the eighteenth international Conf. on Computational Methods in Systems Biology, LNCS. Springer-Verlag, 2020. [ preprint ]
- CMSB 2020b
- Benchmark and Examples as a BIOCHAM notebook
-
Elisabeth Degrand, François Fages, Sylvain Soliman. Graphical Conditions for Rate Independence in Chemical Reaction Networks. In CMSB'20: Proc. of the eighteenth international Conf. on Computational Methods in Systems Biology, volume 12314 of LNCS. Springer-Verlag, 2020. [ preprint ] [ slides ] [ video ]
- ICML workshop CB 2019
- source code
- used for the implementation of
-
Julien Martinelli, Jeremy Grignard, Sylvain Soliman, François Fages. A Statistical Unsupervised Learning Algorithm for Inferring Reaction Networks from Time Series Data. In ICML Workshop on Computational Biology, 2019. [ preprint ]
- CMSB 2019a
- source code
- used for the implementation of
-
Julien Martinelli, Jeremy Grignard, Sylvain Soliman, François Fages. On Inferring Reactions from Data Time Series by a Statistical Learning Greedy Heuristics. In CMSB'19: Proc. of the seventeenth international Conf. on Computational Methods in Systems Biology, volume 11773 of LNCS. Springer-Verlag, 2019. [ preprint ]
- CMSB 2019b
- Biocham-4 notebook of examples of evolved CRNs
- to upload on http://lifeware.inria.fr/biocham4/online/
-
Elisabeth Degrand, Mathieu Hemery, François Fages. On Chemical Reaction Network Design by a Nested Evolution Algorithm. In CMSB'19: Proc. of the seventeenth international Conf. on Computational Methods in Systems Biology, volume 11773 of LNCS. Springer-Verlag, 2019. [ preprint ]
- JTB 2018
- source code
- used for the implementation of
-
Adrien Baudier, François Fages, Sylvain Soliman. Graphical Requirements for Multistationarity in Reaction Networks and their Verification in BioModels. Journal of Theoretical Biology, 459:79–89, 2018. [ preprint ]
- CMSB 2018
- companion BIOCHAM-4 notebooks online
-
Fages, François, Soliman, Sylvain. On Robustness Computation and Optimization in BIOCHAM-4. In CMSB'18: Proc. of the sixteenth international Conf. on Computational Methods in Systems Biology, LNCS. Springer-Verlag, 2018. [ preprint ]
- CMSB 2017a
- companion BIOCHAM-4 notebooks online (zip file of models and notebooks for Biocham v4.0)
-
Fages, François, Le Guludec, Guillaume and Bournez, Olivier, Pouly, Amaury. Strong Turing Completeness of Continuous Chemical Reaction Networks and Compilation of Mixed Analog-Digital Programs. In CMSB'17: Proc. of the fiveteen international Conf. on Computational Methods in Systems Biology, pages 108–127, volume 10545 of LNCS. Springer-Verlag, 2017. [ preprint ]
- CMSB 2017b
- companion BIOCHAM-4 notebooks online (original Python code and examples implementing trace generation and PAC learning, the corresponding biocham commands can also be used directly in Biocham v4)
-
Carcano, Arthur, Fages, François, Soliman, Sylvain. Probably Approximately Correct Learning of Regulatory Networks from Time-Series Data. In CMSB'17: Proc. of the fifteenth international Conf. on Computational Methods in Systems Biology, pages 74–90, volume 10545 of , 2017. [ preprint ]
- TCBB 2018
- Models in Biocham v4.0
- companion notebooks Lotka-Volterra.ipynb, Circadian-Clock.ipynb, p53-Mdm2.ipynb and MAPK.ipynb for Biocham v4.0
-
François Fages, Thierry Martinez, David Rosenblueth, Sylvain Soliman. Influence Networks compared with Reaction Networks: Semantics, Expressivity and Attractors. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 2018. [ preprint ]
- Biosystems 2016
- Files containing models and behavior specification in quantitative temporal logic with BIOCHAM 3.7.5commands for parameter sensitivity and search
-
Pauline Traynard, Céline Feillet, Sylvain Soliman, Franck Delaunay, François Fages. Model-based Investigation of the Circadian Clock and Cell Cycle Coupling in Mouse Embryonic Fibroblasts: Prediction of RevErb-alpha Up-Regulation during Mitosis. Biosystems, 149:59–69, 2016. [ preprint ]
- TCS 2015, CMSB 2012
- Automatic rewriting in SBML of the curated part of biomodels.net from their ODE semantics, and associated graphs.
-
François Fages, Steven Gay, Sylvain Soliman. Inferring Reaction Systems from Ordinary Differential Equations. Theoretical Computer Science, 599:64–78, 2015.
- Constraints 2015, CP 2012
- source code.
-
Faten Nabli, François Fages, Thierry Martinez, Sylvain Soliman. A Boolean Model for Enumerating Minimal Siphons and Traps in Petri-nets. In Proc. of CP'2012, 18th Int. Conf. on Principles and Practice of Constraint Programming, pages 798–814, volume 7514 of LNCS. Springer-Verlag, 2012. [ preprint ] [ slides ] [ poster ]
No longer maintained:
- CellStar image tracker as part of the Yeast Image Toolkit a collective effort for automatic tracking of yeast cells in time-lapse microscopy.
- STSE Spatio-Temporal Simulation environment: a platform facilitating spatial, microscopy-based simulation in biology
- CHRat a modular version of Constraint Handling Rules with ask and tell
- Rules2CP a modelling language for constraint programming
- CLPGUI is a graphical user interface for constraint logic programming.
- GNU Prolog RH is a version of GNU Prolog extended with attributed variables, coroutines and CLP(R) constraints.
- TCLP is a type checker for constraint logic programming in Prolog (source)